Urine-based DNA methylation biomarkers are promising in cancer research
You are here

DNA methylation-based biomarkers have shown potential in disease detection and monitoring. This blog highlights areas in which DNA methylation could support cancer research and examples of biomarkers that are being investigated for various cancer types.
Abnormal DNA methylation is common in cancer
Epigenetic alterations, including aberrant DNA methylation, are among the most common molecular alterations in human cancer. These modifications can occur in early stages of cancer development. Specific genes can also be methylated at various tumor stages and in different cancer types (1).

DNA methylation, unlike RNA and protein alterations, is relatively stable in body fluids and occur in welldefined regions, making it easier to detect by sensitive methylation specific PCR methods (2). DNA methylation signatures are also fairly consistent between genomic DNA from its tissue origins and circulating cell-free DNA (cfDNA), allowing analysis to be extended to almost any body fluid (3,4).
Potential of DNA methylation tests in cancer research
Finding quick, inexpensive yet highly sensitive methods are important in improving cancer detection and management. The value of epigenetic changes as potential biomarker candidates in cancer research is well-established in scientific literature, with many studies associating DNA methylation to clinical parameters. However, many DNA methylation biomarkers still lack large-scale validation, leading to only a few markers being implemented into clinical practice (1,3). Yet several DNA methylation-based in vitro diagnostics (IVD) are available for a wide range of cancer types, including bladder, cervical, colorectal, prostate, glioblastoma, lung breast and liver.
There are at least six broad diagnostic areas in which a DNA methylation cancer test may be combined with traditional screening and medical imaging for better patient outcomes (3):
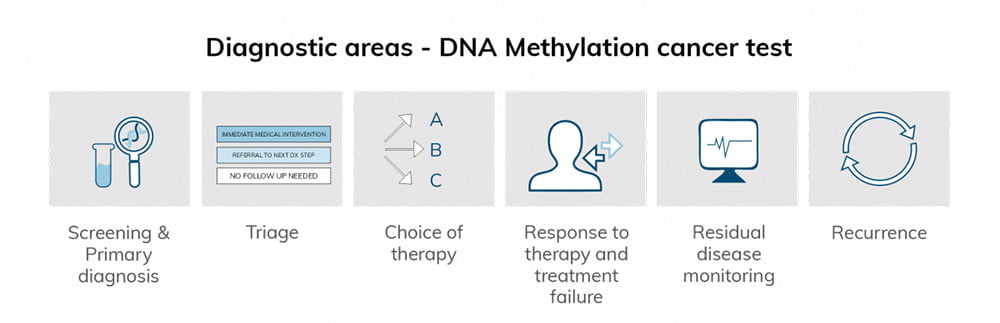
Urine as a liquid biopsy in cancer research
The use of minimally invasive procedures such as liquid biopsies and detection of circulating tumor markers in body fluids, such as blood and urine is gaining interest. The most common liquid biopsy is blood. However, blood contains a relatively high and complex protein repertoire, that can interfere with biomarker measurements. Additionally, blood sampling is relatively invasive, and can pose a risk of infection for both the patient and the caregiver (5).
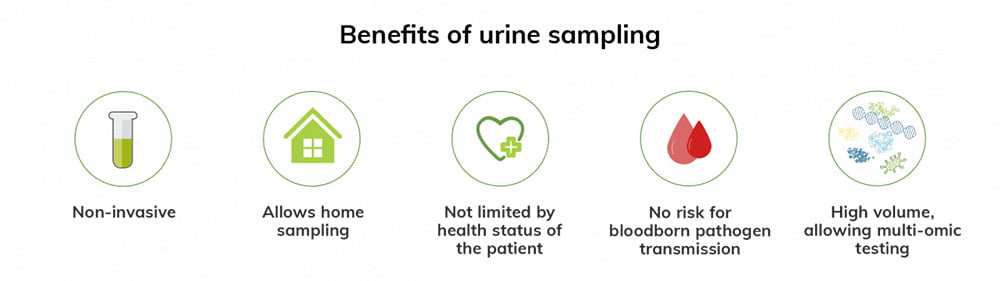
Urine as an alternative biofluid for detecting and monitoring treatment of urological and systemic cancers has shown potential. Urine sampling is easily accessible and available in larger quantities, allows for non-invasive and self-collection and is cost efficient. Sampling is also not limited by the health status of a patient and does not entail any risk of transmission of blood-borne pathogens (5).
Several biomarkers can be found in urine, including circulating molecules such as cfDNA. Urine has also shown to be more a sensitive alternative for early detection or monitoring recurrence of cancers in the genitourinary tract (3).
Examples of common DNA methylation biomarkers in cancer research
Prostate Cancer
The most common (>90%) genetic alteration reported to date in prostate cancer is the epigenetic silencing of the glutathione-S-transferase P1 (GSTP1) gene, as a result of promotor hyper-methylation (6).
Bladder Cancer
Distinct methylation patterns have been identified between non-muscle-invasive bladder cancer (NMIBC) and muscle-invasive bladder cancer (MIBC). For example, methylation of the CDH1, FHIT, LAMC2, RASSF1A, TIMP3, SFRP1, SOX9, PMF1 and RUNX3 genes is associated with poor survival in patients with MIBC (7).
Cervical Cancer
Human Papillomavirus (HPV) is the primary cause of cervical cancer. A fully molecular screen and triage strategy based on primary HPV test and methylation marker detection in the same sample has the potential to increase screening-attendance in hard-to-reach populations and distinguish normal/lowgrade from high-grade underlying cervical disease in high risk-HPV infected women (8).
Future perspectives
DNA methylation-based biomarkers are showing promise in cancer research; these biomarker candidates offer potential to improve screening, and diagnosis but also help better understand recurrence risk and response to therapy. The minimally invasive nature of testing on various body fluid also makes epigenetic-based testing exciting in this field. More work in the area will allow to better understand the scope and value of DNA methylation biomarkers in urine-based cancer research.
Podcast - urine as a liquid biopsy in cancer research
- Constâncio et al. PMID: 32150897
- R Payne et al. PMID: 19459176
- J Locke et al. PMID: 31803237
- Gai et al. PMID: 30634483
- https://novosanis.com/liquid-biopsy-urine-sample-type#5
- Jordaens et al. Urine as an emerging liquid biopsy for prostate cancer biomarkers https://novosanis.com/sites/default/files/poster/pdf/White%20Paper%20PCa...
- Kandimalla et al. PMID: 23628807
- Van Keer et al. PMID: 33846517
